CAS Common Chemistry API in Matlab#
by Anastasia Ramig
CAS Common Chemistry API Documentation (requires registration): https://www.cas.org/services/commonchemistry-api
These recipe examples were tested on April 21, 2022 in MATLAB R2021a.
Attribution: This tutorial uses the CAS Common Chemistry API. Example data shown is licensed under the CC BY-NC 4.0 license.
1. Common Chemistry Record Detail Retrieval#
Information about substances in CAS Common Chemistry can be retrieved using the /detail API and a CAS RN identifier.
Setup API parameters#
detail_base_url = "https://commonchemistry.cas.org/api/detail?";
options = weboptions('Timeout', 30);
casrn1 = "10094-36-7"; % ethyl cyclohexanepropionate
Request data from CAS Common Chemistry Detail API#
casrn1_data = webread(detail_base_url + "cas_rn=" + casrn1, options)
Output:
casrn1_data = struct with fields:
uri: 'substance/pt/10094367'
rn: '10094-36-7'
name: 'Ethyl cyclohexanepropionate'
image: '<svg width="228.6" viewBox="0 0 7620 3716" text-rendering="auto" stroke-width="1" stroke-opacity="1" stroke-miterlimit="10" stroke-linejoin="miter" stroke-linecap="square" stroke-dashoffset="0" stroke-dasharray="none" stroke="black" shape-rendering="auto" image-rendering="auto" height="111.48" font-weight="normal" font-style="normal" font-size="12" font-family="'Dialog'" fill-opacity="1" fill="black" color-rendering="auto" color-interpolation="auto" xmlns="http://www.w3.org/2000/svg"><g><g stroke="white" fill="white"><rect y="0" x="0" width="7620" stroke="none" height="3716"/></g><g transform="translate(32866,32758)" text-rendering="geometricPrecision" stroke-width="44" stroke-linejoin="round" stroke-linecap="round"><line y2="-30850" y1="-31419" x2="-30792" x1="-31777" fill="none"/><line y2="-29715" y1="-30850" x2="-30792" x1="-30792" fill="none"/><line y2="-31419" y1="-30850" x2="-31777" x1="-32762" fill="none"/><line y2="-29146" y1="-29715" x2="-31777" x1="-30792" fill="none"/><line y2="-30850" y1="-29715" x2="-32762" x1="-32762" fill="none"/><line y2="-29715" y1="-29146" x2="-32762" x1="-31777" fill="none"/><line y2="-31376" y1="-30850" x2="-29885" x1="-30792" fill="none"/><line y2="-30850" y1="-31376" x2="-28978" x1="-29885" fill="none"/><line y2="-31376" y1="-30850" x2="-28071" x1="-28978" fill="none"/><line y2="-30960" y1="-31376" x2="-27352" x1="-28071" fill="none"/><line y2="-31376" y1="-30960" x2="-26257" x1="-26976" fill="none"/><line y2="-30850" y1="-31376" x2="-25350" x1="-26257" fill="none"/><line y2="-32202" y1="-31376" x2="-28140" x1="-28140" fill="none"/><line y2="-32202" y1="-31376" x2="-28002" x1="-28002" fill="none"/><text y="-30671" xml:space="preserve" x="-27317" stroke="none" font-size="433.3333" font-family="sans-serif">O</text><text y="-32242" xml:space="preserve" x="-28224" stroke="none" font-size="433.3333" font-family="sans-serif">O</text></g></g></svg>'
inchi: 'InChI=1S/C11H20O2/c1-2-13-11(12)9-8-10-6-4-3-5-7-10/h10H,2-9H2,1H3'
inchiKey: 'InChIKey=NRVPMFHPHGBQLP-UHFFFAOYSA-N'
smile: 'C(CC(OCC)=O)C1CCCCC1'
canonicalSmile: 'O=C(OCC)CCC1CCCCC1'
molecularFormula: 'C<sub>11</sub>H<sub>20</sub>O<sub>2</sub>'
molecularMass: '184.28'
experimentalProperties: [1×1 struct]
propertyCitations: [1×1 struct]
synonyms: {9×1 cell}
replacedRns: []
hasMolfile: 1
Select some specific data#
%% Get Experimental Properties
casrn1_data.experimentalProperties
Output:
ans = struct with fields:
name: 'Boiling Point'
property: '105-113 °C @ Press: 17 Torr'
sourceNumber: 1
%% Get Boiling Point property
casrn1_data.experimentalProperties.property
Output:
ans = '105-113 °C @ Press: 17 Torr'
%% Get InChIKey
casrn1_data.inchiKey
Output:
ans = 'InChIKey=NRVPMFHPHGBQLP-UHFFFAOYSA-N'
%% Get Canonical Smiles
casrn1_data.canonicalSmile
Output:
ans = 'O=C(OCC)CCC1CCCCC1'
2. Common Chemistry API record detail retrieval in a loop#
Setup API parameters#
detail_base_url = "https://commonchemistry.cas.org/api/detail?";
casrn_list = ["10094-36-7", "10031-92-2", "10199-61-8", "10036-21-2", "1019020-13-3"];
Request data for each CAS RN and save to an array#
casrn_data = cell(1,length(casrn_list)); % preallocate
for c = 1:length(casrn_list)
casrn = casrn_list(c);
casrn_data{c} = webread(detail_base_url + "cas_rn=" + casrn);
pause(1); %% add a delay between API calls
end
disp(casrn_data{1, 1}) %% pull out the data for the first value
Output:
uri: 'substance/pt/10094367'
rn: '10094-36-7'
name: 'Ethyl cyclohexanepropionate'
image: '<svg width="228.6" viewBox="0 0 7620 3716" text-rendering="auto" stroke-width="1" stroke-opacity="1" stroke-miterlimit="10" stroke-linejoin="miter" stroke-linecap="square" stroke-dashoffset="0" stroke-dasharray="none" stroke="black" shape-rendering="auto" image-rendering="auto" height="111.48" font-weight="normal" font-style="normal" font-size="12" font-family="'Dialog'" fill-opacity="1" fill="black" color-rendering="auto" color-interpolation="auto" xmlns="http://www.w3.org/2000/svg"><g><g stroke="white" fill="white"><rect y="0" x="0" width="7620" stroke="none" height="3716"/></g><g transform="translate(32866,32758)" text-rendering="geometricPrecision" stroke-width="44" stroke-linejoin="round" stroke-linecap="round"><line y2="-30850" y1="-31419" x2="-30792" x1="-31777" fill="none"/><line y2="-29715" y1="-30850" x2="-30792" x1="-30792" fill="none"/><line y2="-31419" y1="-30850" x2="-31777" x1="-32762" fill="none"/><line y2="-29146" y1="-29715" x2="-31777" x1="-30792" fill="none"/><line y2="-30850" y1="-29715" x2="-32762" x1="-32762" fill="none"/><line y2="-29715" y1="-29146" x2="-32762" x1="-31777" fill="none"/><line y2="-31376" y1="-30850" x2="-29885" x1="-30792" fill="none"/><line y2="-30850" y1="-31376" x2="-28978" x1="-29885" fill="none"/><line y2="-31376" y1="-30850" x2="-28071" x1="-28978" fill="none"/><line y2="-30960" y1="-31376" x2="-27352" x1="-28071" fill="none"/><line y2="-31376" y1="-30960" x2="-26257" x1="-26976" fill="none"/><line y2="-30850" y1="-31376" x2="-25350" x1="-26257" fill="none"/><line y2="-32202" y1="-31376" x2="-28140" x1="-28140" fill="none"/><line y2="-32202" y1="-31376" x2="-28002" x1="-28002" fill="none"/><text y="-30671" xml:space="preserve" x="-27317" stroke="none" font-size="433.3333" font-family="sans-serif">O</text><text y="-32242" xml:space="preserve" x="-28224" stroke="none" font-size="433.3333" font-family="sans-serif">O</text></g></g></svg>'
inchi: 'InChI=1S/C11H20O2/c1-2-13-11(12)9-8-10-6-4-3-5-7-10/h10H,2-9H2,1H3'
inchiKey: 'InChIKey=NRVPMFHPHGBQLP-UHFFFAOYSA-N'
smile: 'C(CC(OCC)=O)C1CCCCC1'
canonicalSmile: 'O=C(OCC)CCC1CCCCC1'
molecularFormula: 'C<sub>11</sub>H<sub>20</sub>O<sub>2</sub>'
molecularMass: '184.28'
experimentalProperties: [1×1 struct]
propertyCitations: [1×1 struct]
synonyms: {9×1 cell}
replacedRns: []
hasMolfile: 1
Select some specific data#
%% Get canonical SMILES
cansmiles = cell(1,length(casrn_data));
for s = 1:length(casrn_data)
smilesnew = string(casrn_data{1, s}.canonicalSmile);
cansmiles{s} = smilesnew;
pause(1);
end
disp(cansmiles)
Output:
Columns 1 through 3
{["O=C(OCC)CCC1CCCCC1"]} {["O=C(C#CCCCCCC)OCC"]} {["O=C(OCC)CN1N=CC=C1"]}
Columns 4 through 5
{["O=C(OCC)C1=CC=CC(=C1)CCC(=O)OCC"]} {["N=C(OCC)C1=CCCCC1"]}
%% Get synonyms
synonyms_list = cell(1,length(casrn_data));
for syn = 1:length(casrn_data)
synonyms_list{syn} = casrn_data{1, syn}.synonyms;
pause(1);
synonyms_list{syn}
end
Output:
ans = 9×1 cell
'Cyclohexanepropanoic acid, …
'Cyclohexanepropionic acid, …
'Ethyl cyclohexanepropionate'
'Ethyl cyclohexylpropanoate'
'Ethyl 3-cyclohexylpropionate'
'Ethyl 3-cyclohexylpropanoate'
'3-Cyclohexylpropionic acid …
'NSC 71463'
'Ethyl 3-cyclohexanepropionate'
ans = 3×1 cell
'2-Nonynoic acid, ethyl ester'
'Ethyl 2-nonynoate'
'NSC 190985'
ans = 5×1 cell
'1<em>H</em>-Pyrazole-1-acet…
'Pyrazole-1-acetic acid, eth…
'Ethyl 1<em>H</em>-pyrazole-…
'Ethyl 1-pyrazoleacetate'
'Ethyl 2-(1<em>H</em>-pyrazo…
ans = 3×1 cell
'Benzenepropanoic acid, 3-(e…
'Hydrocinnamic acid, <em>m</…
'Ethyl 3-(ethoxycarbonyl)ben…
ans = 2×1 cell
'1-Cyclohexene-1-carboximidi…
'Ethyl 1-cyclohexene-1-carbo…
3. Common Chemistry Search#
In addition to the /detail API, the CAS Common Chemistry API has a /search method that allows searching by CAS RN, SMILES, InChI/InChIKey, and name.
Setup API Parameters#
search_base_url = "https://commonchemistry.cas.org/api/search?q=";
options = weboptions('Timeout', 30);
%% InChIKey for Quinine
IK = "InChIKey=LOUPRKONTZGTKE-WZBLMQSHSA-N";
Request data from CAS Common Chemistry Search API#
%% search query
quinine_search_data = webread(search_base_url + IK, options)
Output:
quinine_search_data = struct with fields:
count: 1
results: [1×1 struct]
Note that with the CAS Common Chemistry Search API, only the image data, name, and CAS RN is returned. In order to retrieve the full record, we can combine our search with the related detail API:
%% extract the CAS RN
quinine_rn = quinine_search_data.results.rn;
quinine_rn
Output:
quinine_rn = '130-95-0'
%% get detailed record for quinine
detail_base_url = "https://commonchemistry.cas.org/api/detail?";
quinine_detail_data = webread(detail_base_url + "cas_rn=" + quinine_rn);
quinine_detail_data
Output:
quinine_detail_data = struct with fields:
uri: 'substance/pt/130950'
rn: '130-95-0'
name: 'Quinine'
image: '<svg width="309.3" viewBox="0 0 10310 5592" text-rendering="auto" stroke-width="1" stroke-opacity="1" stroke-miterlimit="10" stroke-linejoin="miter" stroke-linecap="square" stroke-dashoffset="0" stroke-dasharray="none" stroke="black" shape-rendering="auto" image-rendering="auto" height="167.76" font-weight="normal" font-style="normal" font-size="12" font-family="'Dialog'" fill-opacity="1" fill="black" color-rendering="auto" color-interpolation="auto" xmlns="http://www.w3.org/2000/svg"><g><g stroke="white" fill="white"><rect y="0" x="0" width="10310" stroke="none" height="5592"/></g><g transform="translate(32866,32758)" text-rendering="geometricPrecision" stroke-width="44" stroke-linejoin="round" stroke-linecap="round"><line y2="-28559" y1="-28036" x2="-26635" x1="-25742" fill="none"/><line y2="-29819" y1="-28559" x2="-26635" x1="-26635" fill="none"/><line y2="-28036" y1="-28559" x2="-25367" x1="-24474" fill="none"/><line y2="-30451" y1="-29819" x2="-25555" x1="-26635" fill="none"/><line y2="-28559" y1="-29819" x2="-24474" x1="-24474" fill="none"/><line y2="-29504" y1="-28828" x2="-25194" x1="-26005" fill="none"/><line y2="-29819" y1="-30451" x2="-24474" x1="-25555" fill="none"/><line y2="-29082" y1="-28559" x2="-27542" x1="-26635" fill="none"/><line y2="-29819" y1="-30344" x2="-22660" x1="-23567" fill="none"/><line y2="-29700" y1="-30223" x2="-22729" x1="-23636" fill="none"/><line y2="-28779" y1="-29082" x2="-28071" x1="-27542" fill="none"/><line y2="-30703" y1="-30131" x2="-28524" x1="-27542" fill="none"/><line y2="-31850" y1="-30703" x2="-28524" x1="-28524" fill="none"/><line y2="-31705" y1="-30847" x2="-28354" x1="-28354" fill="none"/><line y2="-30131" y1="-30703" x2="-29507" x1="-28524" fill="none"/><line y2="-30131" y1="-30703" x2="-27542" x1="-26560" fill="none"/><line y2="-30347" y1="-30778" x2="-27505" x1="-26768" fill="none"/><line y2="-31850" y1="-32422" x2="-28524" x1="-29507" fill="none"/><line y2="-32312" y1="-31850" x2="-27730" x1="-28524" fill="none"/><line y2="-30703" y1="-30131" x2="-30489" x1="-29507" fill="none"/><line y2="-30778" y1="-30347" x2="-30281" x1="-29544" fill="none"/><line y2="-30703" y1="-31850" x2="-26560" x1="-26560" fill="none"/><line y2="-32422" y1="-31850" x2="-29507" x1="-30489" fill="none"/><line y2="-32205" y1="-31774" x2="-29544" x1="-30281" fill="none"/><line y2="-31850" y1="-32312" x2="-26560" x1="-27354" fill="none"/><line y2="-31760" y1="-32107" x2="-26745" x1="-27340" fill="none"/><line y2="-31850" y1="-30703" x2="-30489" x1="-30489" fill="none"/><line y2="-30275" y1="-30703" x2="-31200" x1="-30489" fill="none"/><line y2="-30541" y1="-30272" x2="-32040" x1="-31575" fill="none"/><polygon stroke-width="1" stroke="none" points=" -24474 -29819 -23602 -30402 -23532 -30284"/><polygon stroke-width="1" points=" -24474 -29819 -23602 -30402 -23532 -30284" fill="none"/><polygon stroke-width="1" stroke="none" points=" -26635 -28559 -26973 -27837 -27092 -27903"/><polygon stroke-width="1" points=" -26635 -28559 -26973 -27837 -27092 -27903" fill="none"/><line y2="-28860" y1="-28796" x2="-25945" x1="-26066" fill="none"/><line y2="-28657" y1="-28611" x2="-25865" x1="-25952" fill="none"/><line y2="-28454" y1="-28427" x2="-25785" x1="-25838" fill="none"/><line y2="-28252" y1="-28242" x2="-25706" x1="-25723" fill="none"/><line y2="-29478" y1="-29530" x2="-25257" x1="-25130" fill="none"/><line y2="-29686" y1="-29727" x2="-25321" x1="-25221" fill="none"/><line y2="-29894" y1="-29924" x2="-25384" x1="-25312" fill="none"/><line y2="-30102" y1="-30121" x2="-25448" x1="-25403" fill="none"/><line y2="-30310" y1="-30317" x2="-25512" x1="-25493" fill="none"/><line y2="-30131" y1="-30128" x2="-27473" x1="-27612" fill="none"/><line y2="-29914" y1="-29912" x2="-27487" x1="-27598" fill="none"/><line y2="-29697" y1="-29695" x2="-27502" x1="-27583" fill="none"/><line y2="-29480" y1="-29479" x2="-27516" x1="-27569" fill="none"/><line y2="-29263" y1="-29263" x2="-27530" x1="-27554" fill="none"/><text y="-28380" xml:space="preserve" x="-28602" stroke="none" font-size="433.3333" font-family="sans-serif">OH</text><text y="-29983" xml:space="preserve" x="-31540" stroke="none" font-size="433.3333" font-family="sans-serif">O</text><text y="-30691" xml:space="preserve" x="-32762" stroke="none" font-size="433.3333" font-family="sans-serif">CH</text><text y="-30602" xml:space="preserve" x="-32185" stroke="none" font-size="313.3333" font-family="sans-serif">3</text><text y="-32242" xml:space="preserve" x="-27695" stroke="none" font-size="433.3333" font-family="sans-serif">N</text><text y="-27747" xml:space="preserve" x="-25708" stroke="none" font-size="433.3333" font-family="sans-serif">N</text><text y="-27473" xml:space="preserve" x="-27311" stroke="none" font-size="433.3333" font-family="sans-serif">H</text><text y="-28600" xml:space="preserve" x="-27695" stroke="none" font-style="italic" font-size="313.3333" font-family="sans-serif">R</text><text y="-28522" xml:space="preserve" x="-26540" stroke="none" font-style="italic" font-size="313.3333" font-family="sans-serif">S</text><text y="-27337" xml:space="preserve" x="-25818" stroke="none" font-style="italic" font-size="313.3333" font-family="sans-serif">S</text><text y="-30573" xml:space="preserve" x="-25708" stroke="none" font-style="italic" font-size="313.3333" font-family="sans-serif">S</text><text y="-29495" xml:space="preserve" x="-24876" stroke="none" font-style="italic" font-size="313.3333" font-family="sans-serif">R</text></g></g></svg>'
inchi: 'InChI=1S/C20H24N2O2/c1-3-13-12-22-9-7-14(13)10-19(22)20(23)16-6-8-21-18-5-4-15(24-2)11-17(16)18/h3-6,8,11,13-14,19-20,23H,1,7,9-10,12H2,2H3/t13-,14-,19-,20+/m0/s1'
inchiKey: 'InChIKey=LOUPRKONTZGTKE-WZBLMQSHSA-N'
smile: '[C@@H](O)(C=1C2=C(C=CC(OC)=C2)N=CC1)[C@]3([N@@]4C[C@H](C=C)[C@H](C3)CC4)[H]'
canonicalSmile: 'OC(C=1C=CN=C2C=CC(OC)=CC21)C3N4CCC(C3)C(C=C)C4'
molecularFormula: 'C<sub>20</sub>H<sub>24</sub>N<sub>2</sub>O<sub>2</sub>'
molecularMass: '324.42'
experimentalProperties: [1×1 struct]
propertyCitations: [1×1 struct]
synonyms: {20×1 cell}
replacedRns: {35×1 cell}
hasMolfile: 1
Handle multiple results#
%% setup search query parameters
search_base_url = "https://commonchemistry.cas.org/api/search?q=";
options = weboptions('Timeout', 30);
%% SMILES for butadiene
smi_bd = "C=CC=C";
%% request data from CAS Common Chemistry Search API
smi_search_data = webread(search_base_url + smi_bd, options);
%% get results count
smi_search_data.count
Output:
ans =
7
%% extract out CAS RNs
smi_casrn_list = {smi_search_data.results.rn};
disp(smi_casrn_list)
Output:
Columns 1 through 5
{'106-99-0'} {'16422-75-6'} {'26952-74-9'} {'29406-96-0'} {'29989-19-3'}
Columns 6 through 7
{'31567-90-5'} {'9003-17-2'}
%% use the detail API to retrieve the full records
detail_base_url = "https://commonchemistry.cas.org/api/detail?";
smi_detail_data = cell(1,length(smi_casrn_list));
for d = 1:length(smi_casrn_list)
casrn = smi_casrn_list(1, d);
smi_detail_data{d} = webread(detail_base_url + "cas_rn=" + string(casrn));
pause(1);
end
%% get some specific data, such as name, from the detail records
names = cell(1,length(smi_detail_data));
for i = 1:length(smi_detail_data)
names{i} = smi_detail_data{1, i}.name;
end
disp(names)
Output:
Columns 1 through 3
{'1,3-Butadiene'} {'Butadiene trimer'} {'Butadiene dimer'}
Column 4
{'1,3-Butadiene, homopolymer, isotactic'}
Column 5
{'1,3-Butadiene-<em>1</em>,<em>1</em>,<em>2</em>,<em>3</em>,<em>4</em>,<em>4</em>-<em>d</e…'}
Columns 6 through 7
{'Syndiotactic polybutadiene'} {'Polybutadiene'}
Handle multiple page results#
The CAS Common Chemistry API returns 50 results per page, and only the first page is returned by default. If the search returns more than 50 results, the offset option can be added to page through and obtain all results:
%% setup search query parameters
search_base_url = "https://commonchemistry.cas.org/api/search?q=";
n = "selen*";
%% get results count for CAS Common Chemistry Search
num_results = webread(search_base_url + n);
%% convert results to an integer
num_results = int16(num_results.count)
Output:
num_results = int16
191
%% request data and save to a cell array in a loop for each page
n_search_data = cell(1,((num_results/50)+1));
for page_idx = 1:((num_results/50)+1)
page_data = webread(search_base_url + n + "&offset=" + string((page_idx-1)*50));
pause(1)
n_search_data{page_idx} = page_data;
end
%% transform the cell array to a table
n_search_data2 = cell2table(n_search_data);
%% create a list of all of the CAS RN values and put it in an array
n_casrn_list = {n_search_data2.n_search_data1.results.rn, n_search_data2.n_search_data2.results.rn, n_search_data2.n_search_data3.results.rn, n_search_data2.n_search_data4.results.rn};
length(n_casrn_list)
Output:
ans =
191
n_casrn_list = n_casrn_list'; %transform
disp(n_casrn_list(1:20))
Output:
{'10025-68-0' }
{'10026-03-6' }
{'10026-23-0' }
{'10101-96-9' }
{'10102-18-8' }
{'10102-23-5' }
{'10112-94-4' }
{'10161-84-9' }
{'10214-40-1' }
{'10236-58-5' }
{'10326-29-1' }
{'10431-47-7' }
{'1049-38-3' }
{'106325-35-3' }
{'1069-66-5' }
{'109428-24-2' }
{'1187-56-0' }
{'1190006-10-0'}
{'1197228-15-1'}
{'12033-59-9' }
% use the CAS RN values and the detail API to obtain the entire record for each
% this will query CAS Common Chem 191 times and take ~ 5 min.
detail_base_url = "https://commonchemistry.cas.org/api/detail?";
n_detail_data = cell(1,length(n_casrn_list));
for casrn = 1:length(n_casrn_list)
n_detail_data{casrn} = webread(detail_base_url + "cas_rn=" + n_casrn_list(casrn), options);
pause(1)
end
%% create a cell array of the molecular mass values
mms = cell(1,length(n_detail_data));
for i = 1:length(n_detail_data)
mms{i} = n_detail_data{1, i}.molecularMass;
end
str_mms = string(mms);
disp(str_mms(1:20)) % view index 1:20
Output:
Columns 1 through 12
"228.83" "220.77" "" "" "" "" "" "300.24" "" "168.05" "" ""
Columns 13 through 20
"" "" "" "241.11" "" "368.25" "265.00" ""
%% remove the empty values
str_mms(strcmp("",str_mms)) = [];
str_mms;
disp(str_mms(1:20))
Output:
Columns 1 through 8
"228.83" "220.77" "300.24" "168.05" "241.11" "368.25" "265.00" "157.92"
Columns 9 through 16
"631.68" "105.99" "154.95" "375.50" "126.96" "559.83" "149.87" "196.11"
Columns 17 through 20
"148.96" "163.00" "231.58" "1174.29"
mms_double = str2double(str_mms);
figure;
x = mms_double;
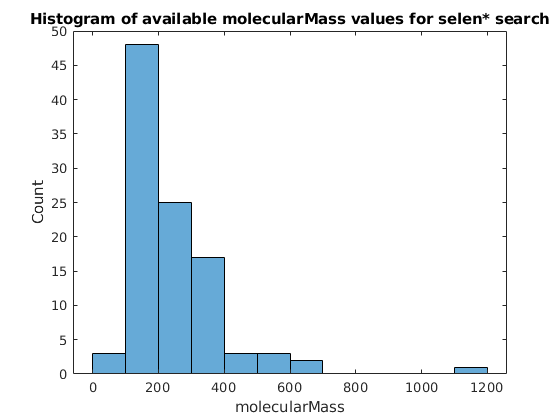
histogram(x)
title("Histogram of available molecularMass values for selen* search");
xlabel("molecularMass");
ylabel("Count");
Output: